NuclearReceptors2018 Oral Communications (1) (8 abstracts)
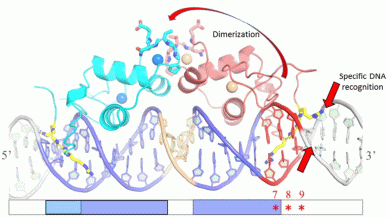
Structural basis of specific DNA recognition by the estrogen-related receptor ERR
Kareeem Mohideen-Abdul , Brice Beinsteiner , Bruno Klaholz , Dino Moras & Isabelle ML Billas
IGBMC, Centre National de la Recherche Scientifique (CNRS), UMR 7104, Institut National de la Santé et de la Recherche Médicale (INSERM) U964, Université de Strasbourg (UdS), Illkirch, 67404, France.
Like other steroid hormone receptors (SHRs), the ERR binds to IR3 response elements (REs). However, the naturally occurring ERR binding sites are composed of single, extended half-sites (ERREs) that are recognized by the homodimeric ERR. In order to unravel the specific structural reasons and the related functional consequences related to the binding of a homodimer to single extended half-site REs, we conducted a series of complementary structural and biophysical experiments and investigated the binding behavior of the isolated DBD to different REs. ERR DBD binds as a monomer to IR3, in contrast with what is observed in the case of ER DBD. However, we also identified target DNA sequences where stable DBD dimer binding is observed. Using a series of chimera and mutant DNA sequences of ERREs and IR3 REs we found the key determinants for the binding of dimeric ERR DBD. Our results suggest that the sequence-directed DNA shape is more important than the exact nucleotide sequence for the binding of ERR DBD to DNA as a dimer. This underlines the importance of the shape-driven DNA readout mechanisms based on minor groove recognition and electrostatic potential. These conclusions may apply not only to ERR, but also to other members of the steroid NR family, such as AR or GR, for which a strong well conserved half-site is followed by a weaker one with a degenerated sequence. We further extended our work to full ERR receptor complexes bound to DNA and present results of structural and functional studies that can be extrapolate to other SHRs. In fact, while the isolated DBD and LBD domains can fulfil their prime functions of transactivation, DNA- and ligand binding, allosteric communication between the different domains is thought to be essential for the cellular function.